News from BOINC Projects
Recent items from the news feeds of various BOINC projects.
[SRBase] base S45 Megaprime
Sightus@CAU, a member of the team Planet 3DNow! found a megaprime for base S45.
The prime 34864*45^679202+1 has 1.122.870 digits digits and entered the T5k PrimePages.
View article · Sat, 9 May 2026 12:21:24 +0000
[SRBase] cable provider down
All services stopped as of May 7, 12:00 UTC+2.
This was the first long major outage since years.
Not a good start for the Pentathlon and certainly not how it was planned.
If the current deadline proves insufficient, it will have to be extended.
Let me know if you have problems with deadlines or other things.
I hope for no more issues on provider side.
Thx for your understanding and support!
View article · Fri, 8 May 2026 01:50:10 +0000
[RakeSearch] Processing of the square # 65 completed
Dear participants, processing of the square # 65 is fully completed! The square:
0 1 2 3 4 5 6 7 8 9 A B
1 2 3 4 5 0 B 6 7 8 9 A
6 B A 9 8 7 4 3 2 1 0 5
4 5 0 1 2 3 8 9 A B 6 7
9 8 7 6 B A 1 0 5 4 3 2
8 7 6 B A 9 2 1 0 5 4 3
5 0 1 2 3 4 7 8 9 A B 6
A 9 8 7 6 B 0 5 4 3 2 1
B A 9 8 7 6 5 4 3 2 1 0
3 4 5 0 1 2 9 A B 6 7 8
2 3 4 5 0 1 A B 6 7 8 9
7 6 B A 9 8 3 2 1 0 5 4
has 25180 diagonal transversals and 513683870 orthogonal mates which puts it on the 35th place of the rating which included at the time of finding 6144 positions. Description and results of other studies can be found on Publications page also.
Now "high part" of spectra of ODLS-12 looks like this (squares # 61 .. 65 marked by magenta):
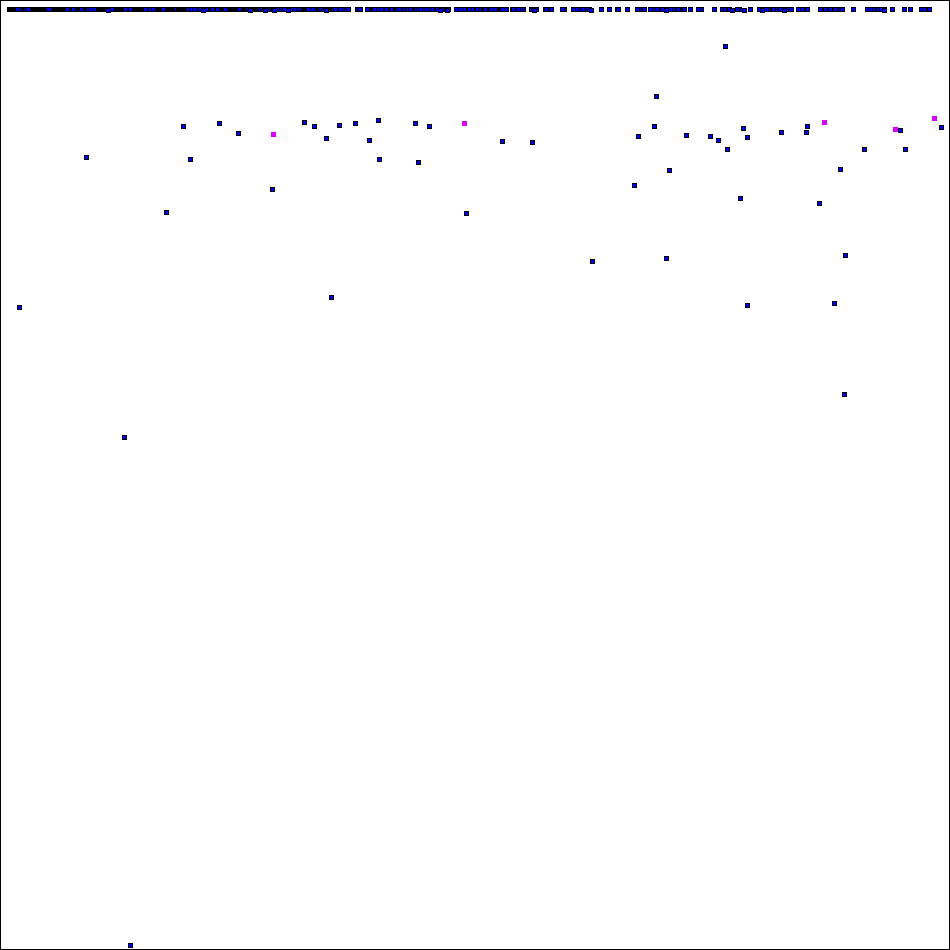
Thank you for project attention, support and donation of CPU time!
View article · Wed, 6 May 2026 08:00:51 +0000
[RakeSearch] Processing of the square # 64 completed
Dear participants, processing of the square # 63 is fully completed! The square:
0 1 2 3 4 5 6 7 8 9 A B
1 2 3 4 5 0 B 6 7 8 9 A
7 6 B A 9 8 3 2 1 0 5 4
2 3 4 5 0 1 A B 6 7 8 9
6 B A 9 8 7 4 3 2 1 0 5
B A 9 8 7 6 5 4 3 2 1 0
9 8 7 6 B A 1 0 5 4 3 2
3 4 5 0 1 2 9 A B 6 7 8
5 0 1 2 3 4 7 8 9 A B 6
A 9 8 7 6 B 0 5 4 3 2 1
8 7 6 B A 9 2 1 0 5 4 3
4 5 0 1 2 3 8 9 A B 6 7
has 25184 diagonal transversals and 465657350 orthogonal mates which puts it on the 53th place of the rating which included at the time of finding 6108 positions.
Thank you for project attention, support and donation of CPU time!
View article · Wed, 6 May 2026 06:03:32 +0000
[RakeSearch] Processing of the square # 63 completed
Dear participants, processing of the square # 63 is fully completed! The square:
0 1 2 3 4 5 6 7 8 9 A B
1 2 3 8 0 9 B 5 4 A 6 7
4 0 1 2 8 7 A B 3 5 9 6
9 A 6 B 5 4 2 8 7 0 1 3
A 6 B 7 9 0 3 4 5 1 2 8
7 5 9 A B 3 0 2 6 8 4 1
6 B 7 5 A 1 8 0 9 2 3 4
8 4 0 1 3 B 9 6 2 7 5 A
B 7 5 9 6 2 4 1 A 3 8 0
5 9 A 6 7 8 1 3 B 4 0 2
3 8 4 0 2 6 5 A 1 B 7 9
2 3 8 4 1 A 7 9 0 6 B 5
has 25194 diagonal transversals and 449892030 orthogonal mates which puts it on the 56th place of the rating which included at the time of finding 6078 positions.
Thank you for project attention, support and donation of CPU time!
View article · Tue, 5 May 2026 21:53:55 +0000
[SRBase] BOINC Pentathlon 2026 - Discipline Javelin Throw
I prefer to bunker average WUs and report them later with
The deadline is 3d + grace_period of 6h and 80 WUs in progress per core.
Only CPU apps are counted.
https://www.seti-germany.de/forum/content/1969-BOINC-Pentathlon-2026-3rd-project-in-the-discipline-Javelin-Throw
View article · Tue, 5 May 2026 06:18:47 +0000
[LHC@home] Downtime Tue 5th of May
Please note that our servers will be unavailable for a while on Tuesday 5th of May until Wednesday 6th due to a change of storage backend.
Hence uploads may fail and other BOINC client requests time out. Many thanks for your understanding and happy crunching!
View article · Mon, 4 May 2026 14:11:07 +0000
[RakeSearch] RakeSearch recovered after technical problems
Dear participants,
thank you for your patience while we were recovering the project. There had been a failure of both SSD discs in a mirror. hoarfrost then did a good job on updating the server and recovering as much data as possible. We had backups of the most important data but not a total backup of everything. I recovered a large part of the forum (sadly excluding the private threads belonging to teams); some particular threads or posts could not be recovered. We hope it does not cause too much inconvenience.
Most of the results you reported before the maintenance have been received, and the credit had been assigned. There may be still some technical glimpses but overall the project is working!
Thank you for participation and the computing power you provide us.
Natalia
View article · Mon, 4 May 2026 09:03:32 +0000
[YAFU] Aliquot sequence 3556280 has terminated!!!
View article · Thu, 30 Apr 2026 19:49:40 +0000
[SRBase] base R48 Megaprime
Jeff Thornton, a member of the team Sicituradastra. found a megaprime for base R48.
The prime 2172*48^668935-1 has 1.124.645 digits digits and entered the T5k PrimePages.
View article · Thu, 30 Apr 2026 12:56:24 +0000
[PrimeGrid] Windows/ARM suppport for GFN apps
PrimeGrid is happy to announce the release of ARM apps for Windows! For many GFN apps that support GPU tasks, we now offer Windows/ARM applications. Just select the CPU tasks in the project preferences page. The list of GFN apps that support Windows/ARM is subject to change. For the most up to date list, see the applications page. Look for "Microsoft Windows running on an ARM 64-bit CPU".
View article · Mon, 27 Apr 2026 19:26:09 +0000
[YAFU] Aliquot sequence 3758388 has terminated!!!
View article · Mon, 27 Apr 2026 18:52:26 +0000
[YAFU] Aliquot sequence 3714280 has terminated!!!
View article · Fri, 24 Apr 2026 19:31:43 +0000
[Gerasim@Home] Gerasim@home:
View article · Fri, 24 Apr 2026 11:46:50 +0000
[SRBase] mfakto-test / fix for 6xxx/7xxx+
Download
Pls run it on windows or linux
win
linux
GPUtype = AUTO /nothing need to be changed
View article · Mon, 20 Apr 2026 20:10:40 +0000
[Amicable Numbers] The search up to 10^21 is finished
After more than 6 years, the BOINC project to find all amicable numbers up to 1021 is finished!
There are 4,810,863 21-digit amicable numbers, and 4,661,814 (96.9%) of them were found by BOINC volunteers. More than 60 million WUs were created and crunched through over the years.
Congratulations everyone and thank you all for your participation!
There are no plans to run the search up to 1022 at the moment - it will take more than 60 years, which is impractical. If, and that's a big IF, a new more efficient exhaustive search method is discovered in the future, then it will make sense to start the project again.
View article · Mon, 20 Apr 2026 08:17:02 +0000
[SRBase] Trial Factoring 76-77bit range started
A new batch of 50k tests is uploading.
The runtime is ~4h40min on a RX5500 XT and decreasing on higher ranges.
The batch name has TF_76-77_ ...
After all 75-76 WUs + resends are done you will see an increase in runtime. GPU in progress will be changed after the challenge.
View article · Sun, 19 Apr 2026 10:14:29 +0000
[Yoyo@home] New App "ECM P2 (20 GB RAM)" in test
I'm currently testing a new app for ECM which requires up to 20 GB of RAM. If you have such a system and want to help testing, please enable the app in your project settings. Validation and credit granting is not yet implemented for this app.
View article · Sat, 18 Apr 2026 22:00:00 +0000
[SRBase] base R42 Megaprime
cjC28bhZdErecrLJfh5u8APsMVa, a member of the team Gridcoin found a megaprime for base R42.
The prime 6964*42^650337-1 has 1.055.663 digits digits and entered the T5k PrimePages.
View article · Sat, 18 Apr 2026 09:22:02 +0000
[SRBase] Formula BOINC 2026-04-16 20:00 UTC - 2026-04-19 20:00 UTC
Good luck everyone and welcome to our new users!
We are nearly done with 75-76bit TF work. The next bitlvl double the runtime starting with 4h40min on an AMD RX5500XT@167M. Keep in mind that AMD 6xxx/7xxx are still not working yet.
View article · Thu, 16 Apr 2026 19:56:42 +0000
[NFS@home] New Cunningham project record
The recently completed factorization of 12,311- into 155-digit and 181-digit prime factors set a new Cunningham project record for the largest penultimate prime factor. Congratulations to all of the project volunteers!
View article · Wed, 8 Apr 2026 23:05:00 +0000
[Minecraft@Home] Diamond Badges and Progress Update
Hello,
As promised, badges have been added for the diamond ore vein project, there are currently 3 badges, the first is given to all diavein project contributors, second is reserved for the top 10% of credit holders and third is reserved for the top 5% this gets updated every 24 hours so it is possible to loose a badge if your credit amount no longer meets those requirements.
As for a project progress update, things are going well, we have generated the next 65k workunits, and the largest vein we've found so far is 58 vein. We're gunning for the first 60+

Happy Crunching!
- scriptline
View article · Wed, 8 Apr 2026 20:59:55 +0000
[YAFU] Aliquot sequence 3678234 has terminated!!!
Aliquot sequence 3678234 has terminated!!!
View article · Tue, 7 Apr 2026 18:35:25 +0000
Copyright © 2026 University of California.
Permission is granted to copy, distribute and/or modify this document
under the terms of the GNU Free Documentation License,
Version 1.2 or any later version published by the Free Software Foundation.